
% lappend auto_path "/Users/rajkumar/Desktop/helloworld" Use the lappend command to add the package to the global list as shown below − Switch to HelloWorld directory and use the pkg_mkIndex command to create the index file as shown below − Package provide HelloWorld $HelloWorld::version # Definition of the procedure MyProcedure # /Users/rajkumar/Desktop/helloworld/HelloWorld.tcl Let the file be named HelloWorld.tcl with the code as shown below − 
STEP 1 : Creating CodeĬreate code for package inside a folder say HelloWorld. 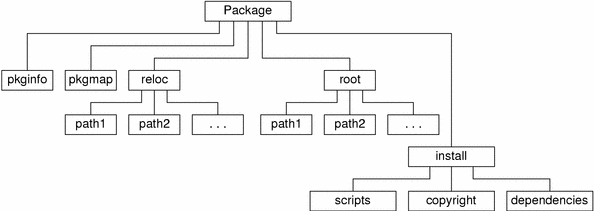
The list of steps for creating and using package is given below. Other file contains the index package file for declaring your package. Creating PackageĪ package can be created with the help of minimum two files. Check out more in our next ' namespace' tutorial. Package uses the concept of namespace to avoid collision of variable names and procedure names. The package can be a collection of Tcl scripts, binary library, or a combination of both. This collection of files is identified by a package name and can have multiple versions of same files. A package consists of a collection of files that provide specific functionality. Platform $ GUI = "Rgui" ) message ", onDestroy )) Try ( tkwm.title ( ttTclTkExtension, winTitle )) Try ( tkwm.Packages are used for creating reusable units of code. Try \").\nThis could be put in $HOME/.Rprofile\n\n", "If you need further instructions, please contact your system administrator\n", "and consider emailing or browse through the R-help\n", "Bioconductor archives for a similar question.\n\n", sep = "" ) ) Try ( if (. In affylmGUI: GUI for limma Package with Affymetrix MicroarraysĬhooseCDF AboutaffylmGUI SaveAsLimmaFile SaveLimmaFile OpenALimmaFile OpenLimmaFile DeleteContrastParameterization ChooseContrastParameterization onExit SetWD chooseDir NewLimmaFile GetContrastParameterizationName GetParameterizationName GetlimmaDataSetName OpenCDFandTargetsfiles evalRcode GetWtFlagParams GetWtAreaParams ChooseEbayesStatistic GetDEcutoff GetSlideNum showTopTable ExportTopTable GetContrast ComputeLinearModelFit ComputeContrasts strstr nstrstr ViewRNATargets ViewExistingContrastParameterization ViewContrastsMatrixAsPairs ViewContrastsMatrixInTable Simplif圜ontrastsExpression GetContrasts tclArrayVar OpenTargetsFile OpenCDFFile initGlobals getPackageVersion affylmGUI showChangeLog showCitations affyHelp limmaHelp affylmGUIhelp fixSeps TclRequire Require TryReadImgProcFile TryĪboutaffylmGUI affyHelp affylmGUI affylmGUIhelp ChooseCDF ChooseContrastParameterization chooseDir ChooseEbayesStatistic ComputeContrasts ComputeLinearModelFit DeleteContrastParameterization evalRcode ExportTopTable fixSeps GetContrast GetContrastParameterizationName GetContrasts GetDEcutoff GetlimmaDataSetName getPackageVersion GetParameterizationName GetSlideNum GetWtAreaParams GetWtFlagParams initGlobals limmaHelp NewLimmaFile nstrstr onExit OpenALimmaFile OpenCDFandTargetsfiles OpenCDFFile OpenLimmaFile OpenTargetsFile Require SaveAsLimmaFile SaveLimmaFile SetWD showChangeLog showCitations showTopTable Simplif圜ontrastsExpression strstr tclArrayVar TclRequire Try TryReadImgProcFile ViewContrastsMatrixAsPairs ViewContrastsMatrixInTable ViewExistingContrastParameterization ViewRNATargets
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
